Notice: This Wiki is now read only and edits are no longer possible. Please see: https://gitlab.eclipse.org/eclipsefdn/helpdesk/-/wikis/Wiki-shutdown-plan for the plan.
STEM Model Creator/Tutorials
Contents
Tutorial 1: Create a textbook SEIR model by extending SIR
Getting Started
This tutorial shows how to create a new SEIR Disease Model by extending the existing SIR Disease Model using the STEM Model Generator, Visual Editor, and Expression Language - all without writing a line of Java code. STEM already has a built-in SEIR model, so this is more a how-to demo for creating new disease models.
Before you begin, you should understand how compartment models work in STEM. Also, please look over the STEM Model Creator and STEM Expression Language documents.
Create Model Project and Disease Model
- Launch the STEM application
- Open the Designer perspective
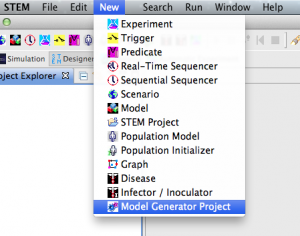
- From the New menu, select Model Generator Project
- Choose Create and configure a new STEM Model Package
- Click Next
- On the Model Package page, enter the following values:
- Package Name: Demo
- Package Prefix: com.example.diseasemodels
- Click Add Model to add the disease model
- On the Model Properties page, enter these values:
- Model Name: MySEIR
- Model Type: Disease Model
- Parent Model: Select SIR from the list
- Click Finish
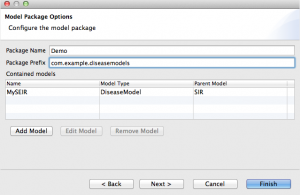
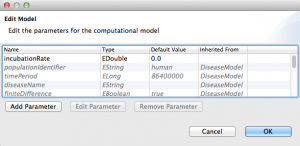
- At this point, your package properties should look like this:
- Click Finish to run the Model Generator and create your model project
- This step may take 1-2 minutes to complete
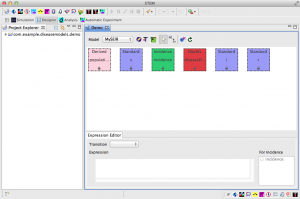
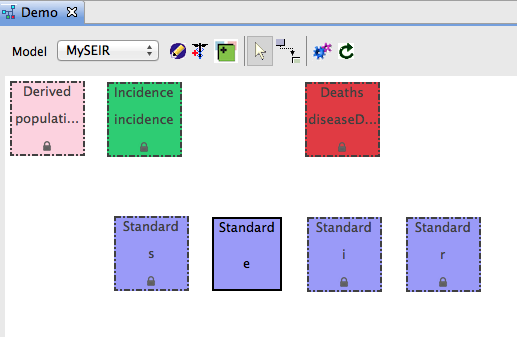
- After the Model Generator finishes, the Visual Editor should automatically open
Add New Compartment
Now that your model project is created, it's time to add new parameters and compartments. First, let's create a new compartment.
We're creating an S-E-I-R model, which is made up of compartments S, E, I, and R. Since we extended the built-in S-I-R model, we need to add a new E compartment.
- In the Visual Editor, click the Add Compartment button
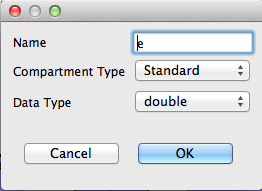
- In the Add Compartment dialog, enter the following values
- Name: e (make sure this is lower case)
- Compartment Type: Standard
- Data Type: double
- Click Finish
- A new e box now appears in the diagram
- You may want to rearrange or resize the compartment boxes so that the S, E, I, and R boxes are in order (easier when connecting the compartments)
That's it. You've added a new compartment to your model. This is now a S-E-I-R model.
Add New Model Parameter
In addition to inheriting compartments from the parent SIR model, the new model also inherits the model (or rate) parameters from SIR. Inherited parameters include Transmission Rate (transmissionRate), Recovery Rate (recoveryRate), Immunity Loss Rate, etc.
We already added the new Exposed state to the disease model. In the textbook SEIR example, the rate controlling movement from the E to I compartments is the Incubation Rate. The next step is to add the incubationRate model parameter to the MySEIR model.
- In the Visual Editor, click the Edit Model button
- In the Edit Model dialog, click the Add Parameter button
- In the Edit Parameter dialog, enter the following values:
- Name: incubationRate
- Display Name: Incubation Rate
- Data Type: double (default)
- Default Value: 0.0 (default)
- Optionally, add a minValue constraint of 0.0
- Click OK to add the parameter
- Click OK again to apply the changes
Regenerate the Model Project
Changing a model's parameters or compartments requires a Model Regeneration. This operations rebuilds several of the key Java files that make up the structure of your model project. For more information, see the section on Model Regeneration.
To run the Model Generator:
- In the Visual Editor, save the MySEIR model
- Click the Model Generator button in the Visual Editor Toolbar
- If prompted, save the model file
- In the Model Generator wizard, immediately click Finish
- Wait while the model is regenerated
- This may take up to a minute to complete
After the Model Generator wizard closes, you're ready to implement the model's transitions and expressions.
Create Transitions and Expressions
Now that the initial setup is finished, you're now ready for the fun part - creating the Transitions that describe the movement of a population between compartments in the disease model.
Transitions have four pieces:
- The source compartment
- The target compartment
- The equation that describes the flow rate out of the source compartment and into the target compartment per time step
- The list of incidences the transition counts toward
With the Visual Editor, you can easily draw the transitions as directed graph edges between a source and target compartment then use the STEM Expression Editor to enter the rate equations.
Draw Transitions
For the MySEIR model, we will create transitions between these compartments
Source Target Transition s e S → E e i E → I i r I → R i diseaseDeaths I → diseaseDeaths r s R → S
To draw transitions:
- In the Visual Editor, select the Draw Transition Tool button
- To create the S → E transition:
- Select the S compartment as the source. Click once on the s compartment box
- Select the E compartment as the target. Move the mouse to and click once on the e compartment box
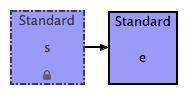
- You should now see a directed connector between the s and e compartment boxes
- Repeat for the other transitions listed above
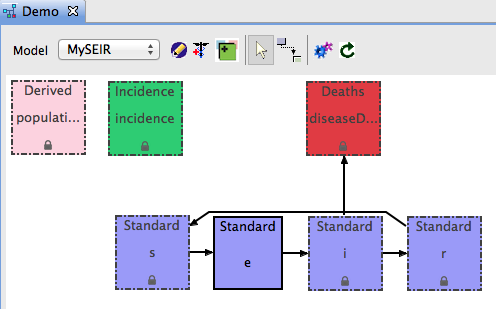
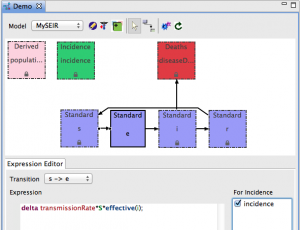
- After you create all five transitions, your diagram should look like this:
Enter Rate Equations
For our MySEIR model, the next step is to enter the equations for the five transitions we created in the previous step.
Using the Expression Editor , you can easily enter the equations that represent the state changes in your model as STEM Expressions. For more information, please read about the STEM Expression Language and the Expression Editor.
- In the Visual Editor, select the Chooser tool in the toolbar
- Select connector edge between the s and e compartment boxes
- In the Expression Editor , enter the equation expression
- Note: The final statement starts with the word delta and end with a semicolon ( ; )
- For S → E, enter:
-
delta transmissionRate*S*effective(i);
-
- Tip You can use auto-completing expression assistance as you enter your expression. Just type Control - S as you type for the assistance dialog!
- For S → E, select the box next to incidence under For Incidence
- This tells the STEM simulation engine to count this transition as part of the disease incidence
- Repeat these steps for the other transitions in the model, entering these expressions for the transition:
Transition Expression Incidences to select S → E delta transmissionRate*S*effective(i);incidence E → I delta incubationRate*E;none I → R delta recoveryRate*I;none I → diseaseDeaths delta infectiousMortalityRate*I;none R → S delta immunityLossRate*R;none
Save your Model
Before you continue, be sure to Save the model. This will tell STEM to generate the Java code that implements your disease model and install it into the STEM application.
To save the open model in the Visual Editor, either do Control - S ( Command - S on Mac OS X) or open the File menu and select Save.
Summary
That's it! You've now created a simple, textbook S-E-I-R disease model using the STEM Model Creator. In the next section/tutorial, you'll learn how to use the new model in a STEM simulation.
Tutorial 2: Run a Disease Simulation using the MySEIR model
Getting Started
In the previous tutorial, you used the STEM Model Generator to create and build a fully functional SEIR disease model in STEM. You're now ready to run your new MySEIR disease model in STEM. If you're new to STEM, at this point you may want to look through the STEM Scenario guide to understand how STEM scenarios, models, and simulations work.
To simplify this tutorial, we've provided a simple demo scenario project with pre-built scenarios, geographic models, infectors, innoculators, and sequencers. In this tutorial, we'll import the project into STEM, create a new disease instance of your MySEIR disease model, add the disease to a geographic model, and run a STEM simulation.
Download and Import Demo Project & Scenario
The first step is to download the VisualEditorDemo project and import it into STEM
- Download the Visual Editor Demo project zip file
- In the STEM application, open the File menu and select Import
- In the Import Wizard, under General, select Existing projects into Workspace
- Click Next
- Click the radio button next to Select Archive File
- Click Browse and navigate to the location you downloaded ZIP file
- Select VisualEditorDemo.zip and click Open
- Verify VisualEditorDemo is listed under Projects
- Click Finish
Create MySEIR Disease
Next, we want to create an instance of our new MySEIR disease model.
- In STEM, open the New menu and select Disease
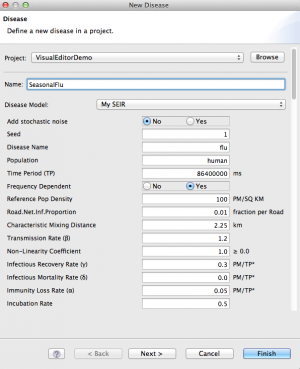
- In the New Disease wizard, make sure VisualEditorDemo is the selected project
- For Name, enter SeasonalFlu
- From the Disease Model drop-down, choose MySEIR (probably at the bottom of the list)
- For Disease Name, enter flu (lower case)
- Enter appropriate rate parameter values. For this demo, here are some suggestions:
- Transmission Rate: 1.2
- Infectious Recovery Rate: 0.3
- Immunity Loss Rate: 0.05
- Incubation Rate: 0.5
- All others: defaults
- Click Finish to create the new disease
Add Disease to Geographic Model
Next, we'll add the new Disease to the pre-built geographic model.
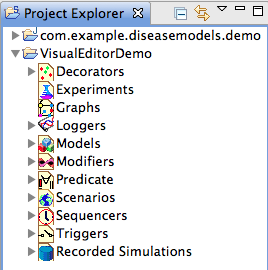
- In the STEM Project Explorer, expand the VisualEditorDemo project
- Expand the Models folder. Double-click on CubaDisease.model
- The CubaDisease.model editor should open
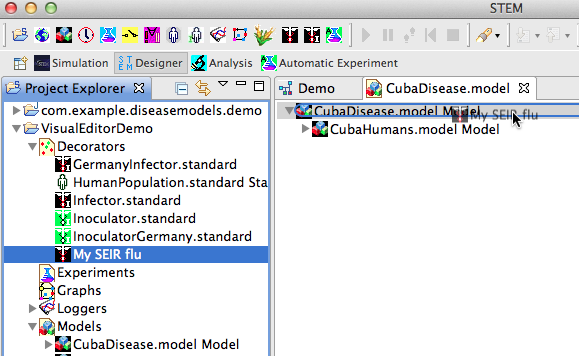
- In Project Explorer, expand the Decorators folder
- Select My SEIR flu, drag and drop into the CubaDisease.model editor root
- Save and Close the CubaDisease.model editor
Launch the Scenario as a STEM Simulation
You're finally ready to see the results of all the work. You will now launch the pre-built scenario as a STEM simulation and observe the epidemic curve as a time series chart.
- In the Project Explorer, expand the Scenarios folder
- Right Click on CubaDemo.scenario and select Step
- On the Graph Map, double click on the Cuba polygon to open a Time Series tracker
- Under Simulation, click the Run Button
 to start the simulation
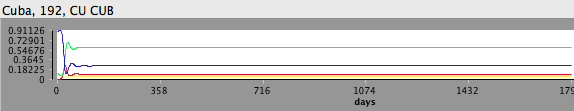
to start the simulation - Wait while the simulation runs, watching the Time Series view of the epidemic curve
- Your time series plot should look like this:
- When the simulation finishes, you can click the Stop Button

Summary
You're finished! You've successfully authored a simple, textbook SEIR disease model, created a disease instance in STEM, and run a simple spatiotemporal simulation. Obviously this is not a real-world example, but it's a good introduction to using the STEM Model Creator, Visual Editor, and Expression Language to easily author compartmental models of disease.
In the next tutorial, you will build on this existing model to add more complicated features, such as sinusoidal and temperature forcing to create a seasonal variation in your disease model.
Tutorial 3: Add seasonal modulation to the SEIR model
Getting Started
In this tutorial, we are going to extend the MySEIR disease model previously created to add a seasonal modulation to transmission. We will create two different modulations - one using a sinusoidal function and then one that uses the STEM Earth Science data to be temperature-dependent.
Sinusoidal
In this example, we'll add a Modulation Period parameter to the MySEIR then use the built-in sin(...) function to apply a sinusoidal modulation to the transmission rate.
If you've previously closed the Visual Editor, reopen the MySEIR model
- In the Project Explorer expand com.example.diseasemodels.demo project
- Expand the model folder
- Double click on demo.metamodel to open the Visual Editor
- Add the Modulation Period parameter (Adding a model parameter)
- Name: modulationPeriod
- Display Name: Modulation Period
- Data Type: double
- Default Value: 365.25 (one year)
- Add a minValue constraint of 0.0
- Remember to run the Model Regenerator
- Select the S → E transition
- In the Expression Editor
- First add a seasonalModulation local variable
-
seasonalModulation = 1+sin(2*3.14*t/modulationPeriod);
-
- Then add the seasonalModulation to the expression delta:
-
delta transmissionRate*S*effective(i)*seasonalModulation/2
-
- First add a seasonalModulation local variable
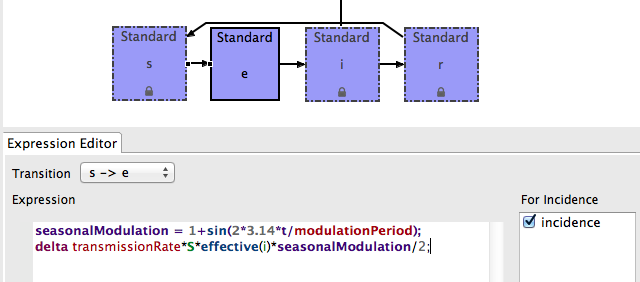
- Final expression should look like:
- Save the MySEIR model
- Now re-run the CubaDemo.scenario simulation to see the change in the Time Series graph
Temperature-dependent
Next you're going to change the seasonal modulation from a sinusoidal function to be temperature-dependent. To get temperature data, you'll use the temperature() function that comes with the STEM Expression Language. The temperature() function uses the STEM Earth Science data applied to each region in your geographic model.
To access temperature data, you must first:
- Install the Earth Science Add-on into STEM
- Add the Earth Science data labels to your geographic model
- In our VisualEditorDemo project, this the data is already added to the Germany model
Next change the expression to add the temperature() function
- In the Visual Editor, select the MySEIR model
- Select the S → E transition
- In the Expression Editor, change the expression to:
-
seasonalModulation = max(temperature()/10,0); -
delta transmissionRate*S*effective(i)*seasonalModulation;
-
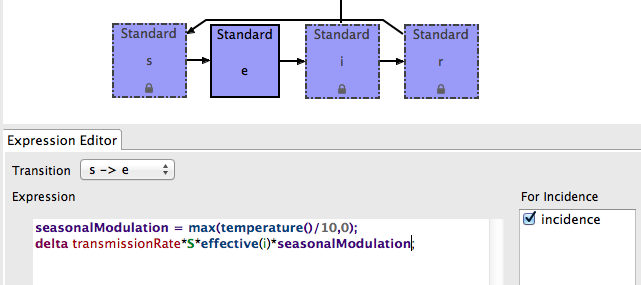
- This is what the expression should look like:
- Save the MySEIR model
- In the Project Explorer, expand the VisualEditorDemo project
- Under models, open GermanyDisease.model
- Drag and drop the MySEIR flu disease into the GermanyDisease.model root
- Save and Close the GermanyDisease.model editor
- Next, under Scenarios, right click on GermanyDemo.scenario and select Run
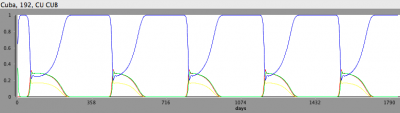
- In the Simulation perspective, double click on one of the Germany regions to add a Time Series tracker
- You may need to restart the scenario after this
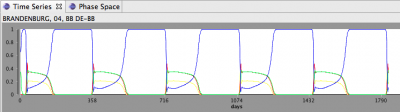
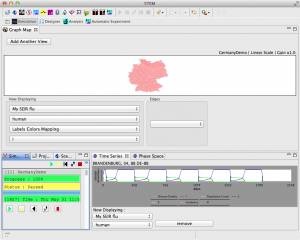
- After the simulation finishes, you should see a Time Series that looks like this:
Summary
You've now successfully modified a textbook SEIR model to add a simple seasonal modulation. At this point, you can now begin exploring how to use the STEM Model Creator and Expression Language to tackle more sophisticated disease models.
More tutorials will be coming soon!