Notice: this Wiki will be going read only early in 2024 and edits will no longer be possible. Please see: https://gitlab.eclipse.org/eclipsefdn/helpdesk/-/wikis/Wiki-shutdown-plan for the plan.
STEM Avian Influenza Model
Background
Several outbreaks of avian influenza have occurred in the past, including the SARS outbreak in 2002, H5N1 in 1997 and H7N9 in 2013 -- all originating in China. This STEM model captures the transmission of the virus between (domesticated) birds, from birds to humans (e.g. at market places) and optionally human-to-human. Mixing of infectious birds between neighboring regions is controlled using a separate parameter from the human mixing parameter.
Model
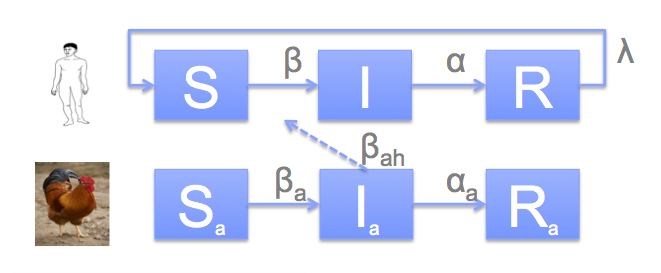
The model is uses an SIR model for humans as well as birds, as well as an indirect transmission pathway between infected birds and susceptible humans:
Below are the differential equations that explain the model:
When the model contains temperature earth science data, a seasonal effect in transmission is included in the mass action term. You can download an example STEM scenario of an Avian Influenza outbreak here: [1]
These are the parameters:
Avian Characteristic Mixing Distance. Controls mixing of infected birds across neighboring regions. Higher values means more mixing
Avian Recovery Rate. &lambda&a Rate of recovery for infected birds (1/time period)
Avian to Avian Transmission Rate. Transmission rate between birds
Avian to Human Transmission Rate. β Transmission rate between birds and humans
Characteristic Mixing Distance. Mixing of infected humans across neighboring regions. Higher values mean more mixing
Human Immunity Loss Rate. Immunity loss rate for humans (1/time period)
Human Recovery Rate. Human recovery rate (1/time period)
Human to Human Transmission Rate. Transmission rate between humans
Temperature Factor. The factor controlling linear scaling of the transmission with temperature
Vector Population Identifier. The population identifier for the avian population